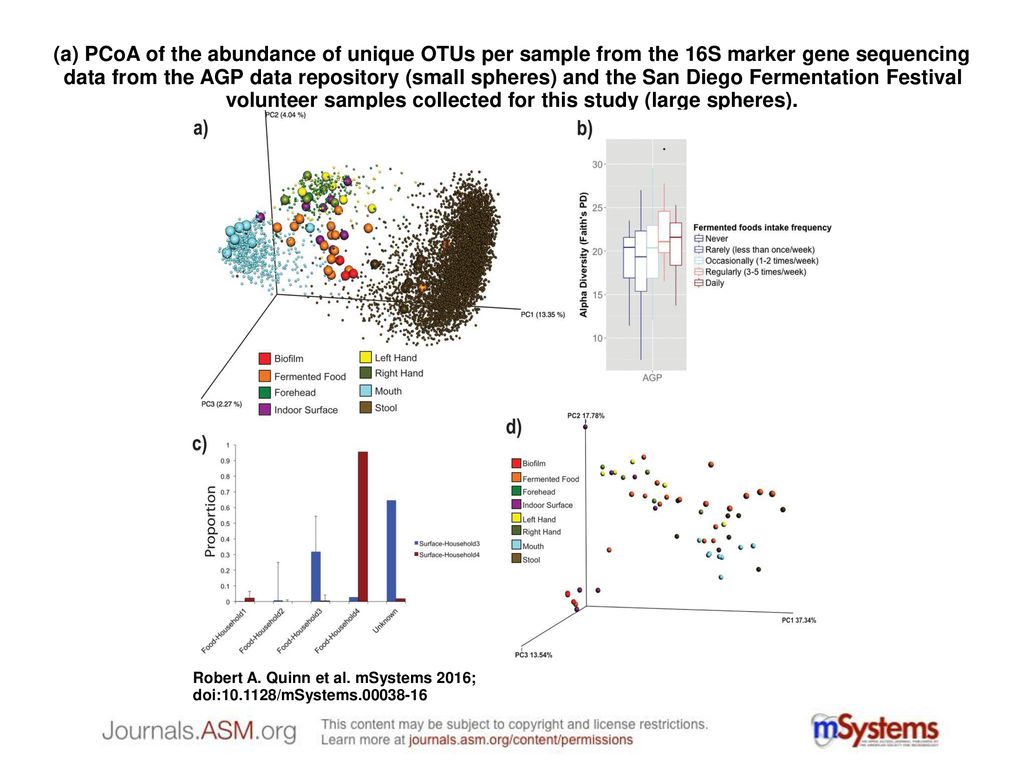
a) PCoA of the abundance of unique OTUs per sample from the 16S marker gene sequencing data from the AGP data repository (small spheres) and the San Diego. - ppt download

Gene Expression Changes and Community Turnover Differentially Shape the Global Ocean Metatranscriptome. - Abstract - Europe PMC

MUSiCC: a marker genes based framework for metagenomic normalization and accurate profiling of gene abundances in the microbiome | Genome Biology | Full Text

Inferring gene regulation from stochastic transcriptional variation across single cells at steady state | PNAS

Abundance of microbial CO2-fixing genes during the late rice season in a long-term management paddy field amended with straw and straw-derived biochar
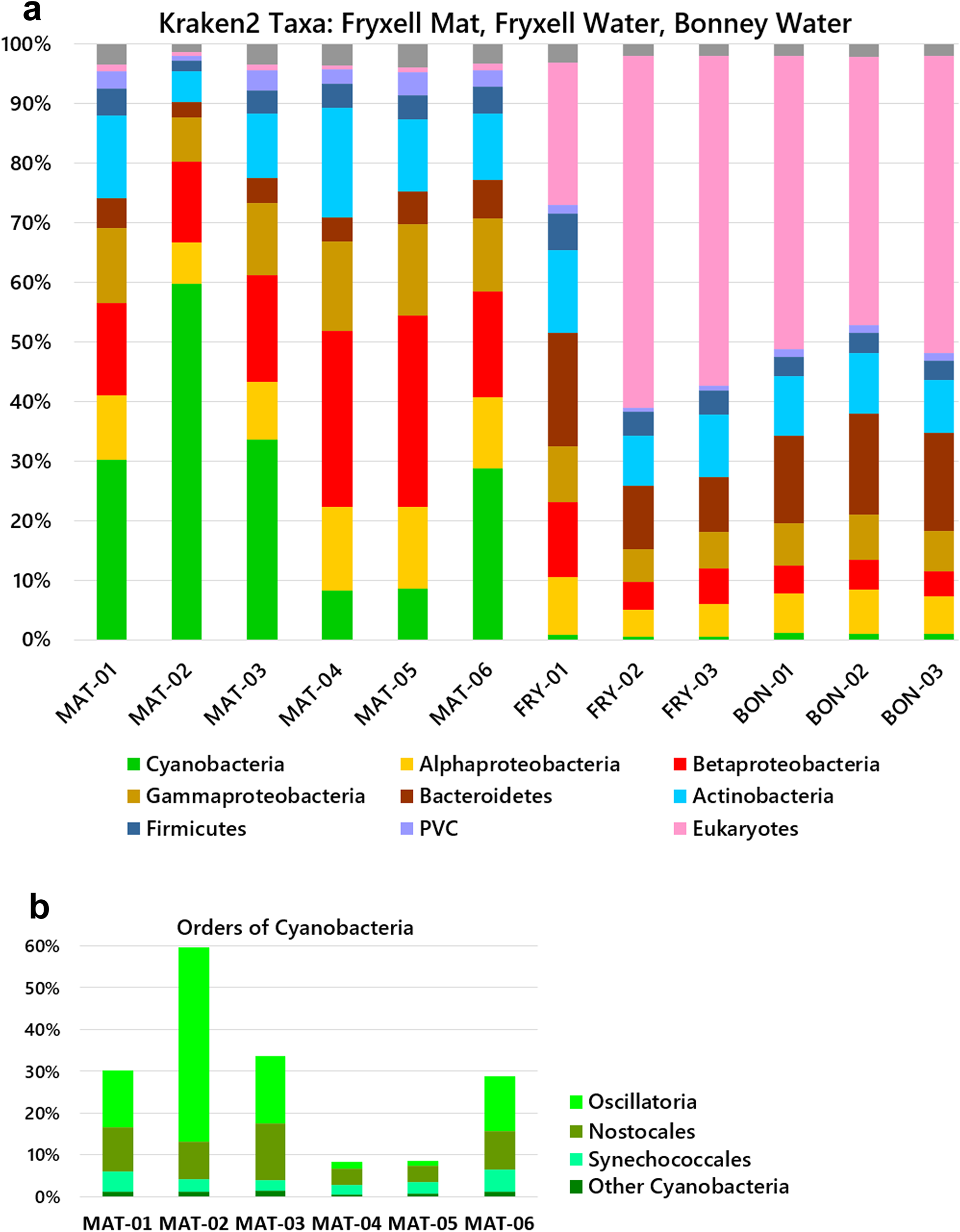
Antibiotic resistance genes and taxa analysis from mat and planktonic microbiomes of Antarctic perennial ice-covered Lake Fryxell and Lake Bonney | Antarctic Science | Cambridge Core

HUMAnN calde-specific marker genes, nucleotide, protein database note available on .org server - usegalaxy.org support - Galaxy Community Help

A neural network‐based framework to understand the type 2 diabetes‐related alteration of the human gut microbiome - Guo - 2022 - iMeta - Wiley Online Library
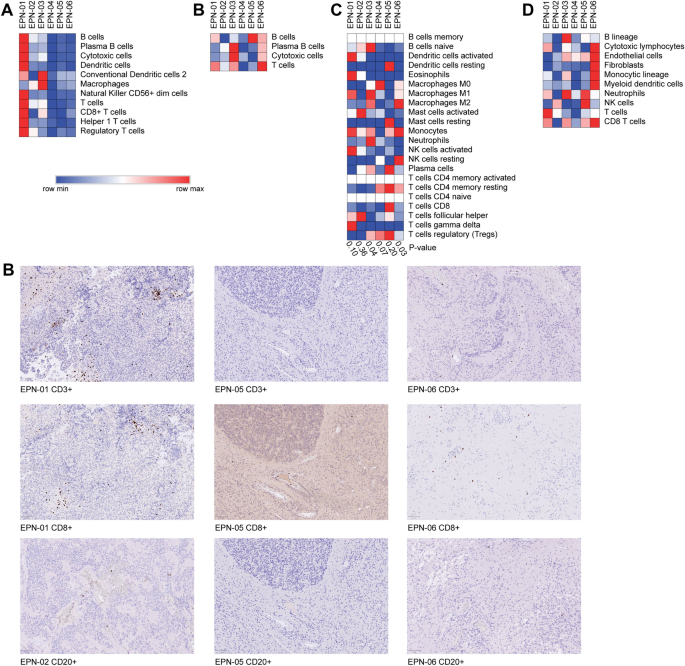
Characterizing the tumor immune microenvironment of ependymomas using targeted gene expression profiles and RNA sequencing | Cancer Immunology, Immunotherapy

Neighbor-specific gene expression revealed from physically interacting cells during mouse embryonic development | PNAS

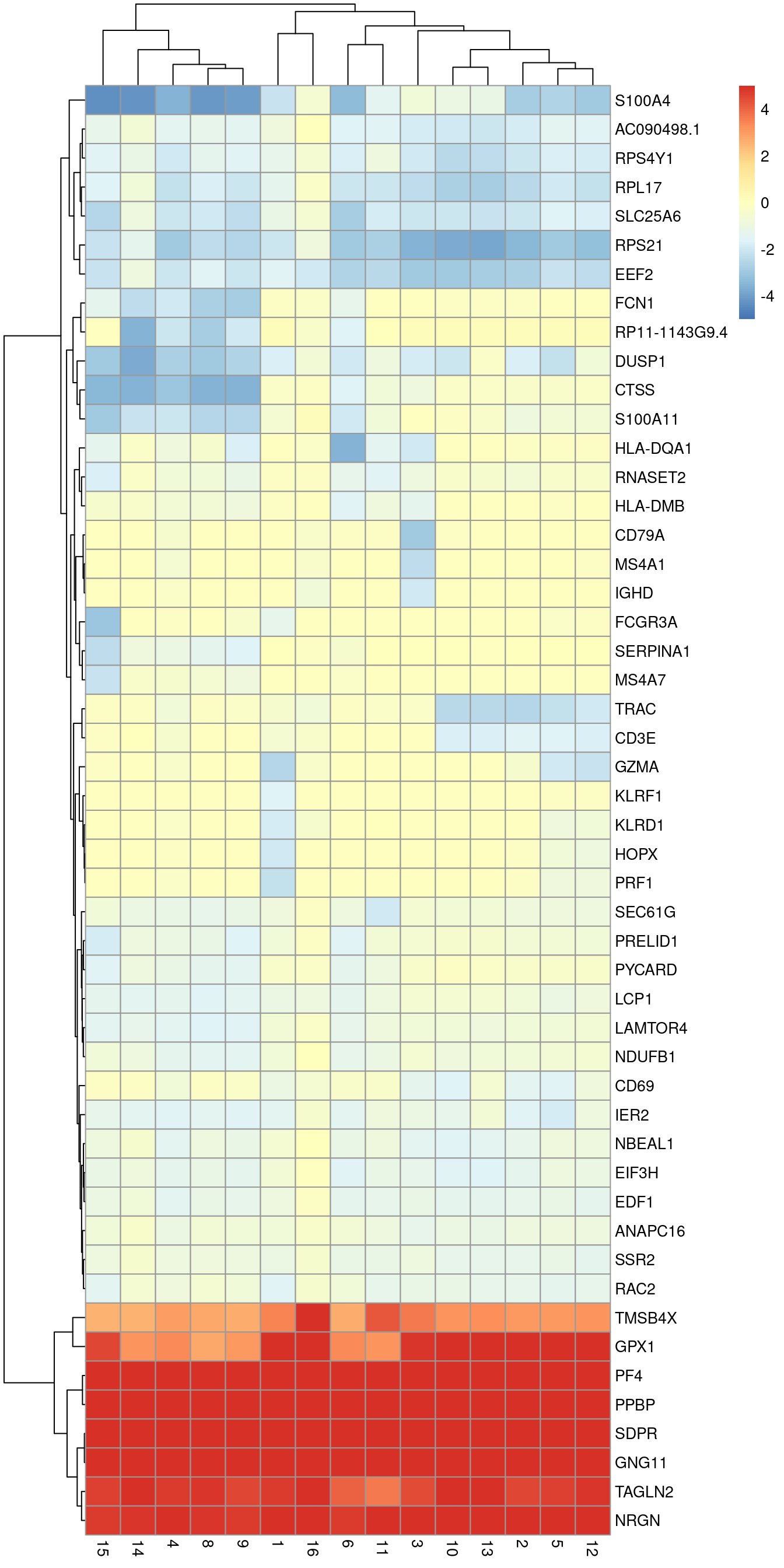
Michael VanInsberghe on X: "Cluster analysis and pseudotime ordering of these cells revealed not only the expected change in abundances of known cell-cycle marker genes, but also that the frequency of certain
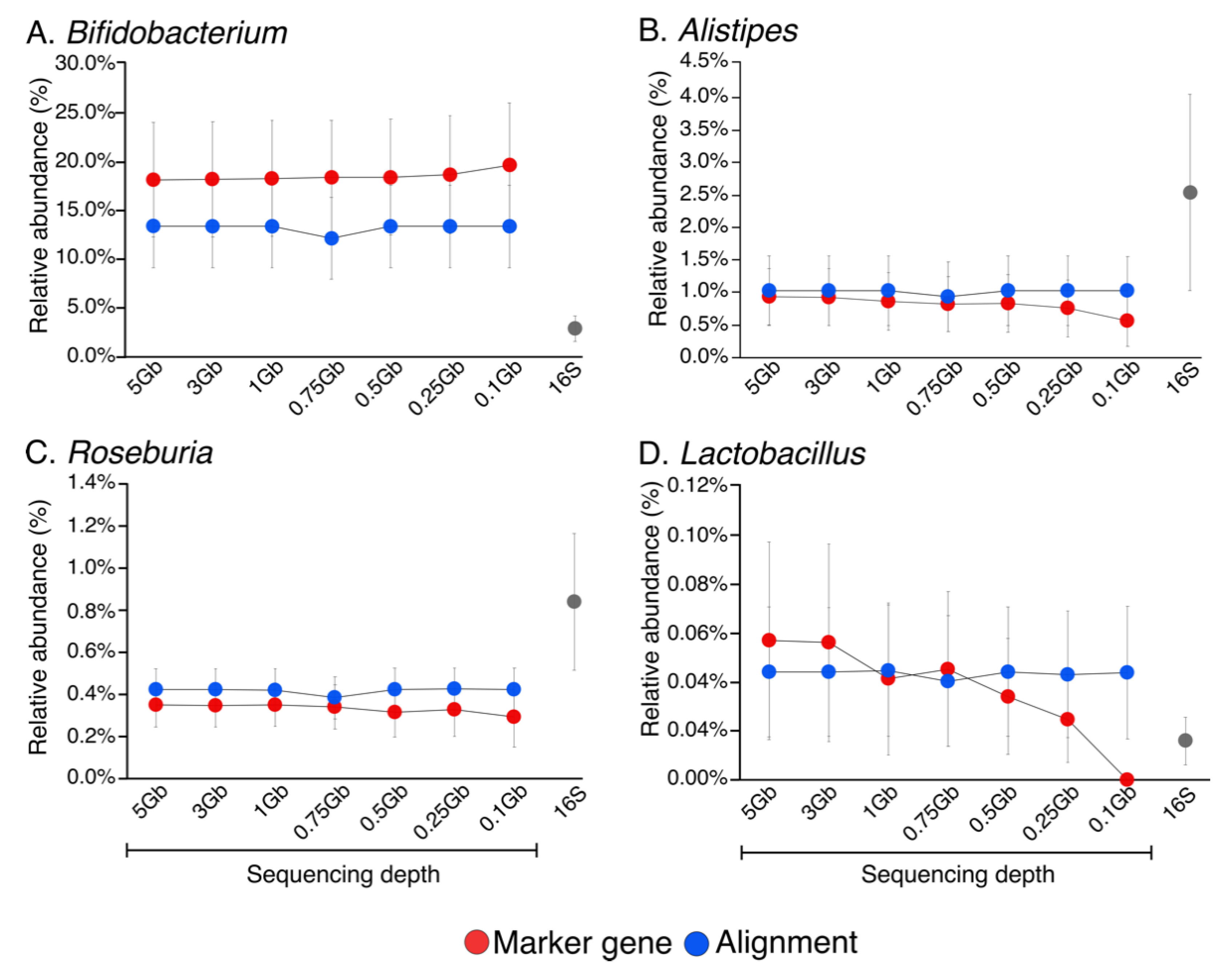
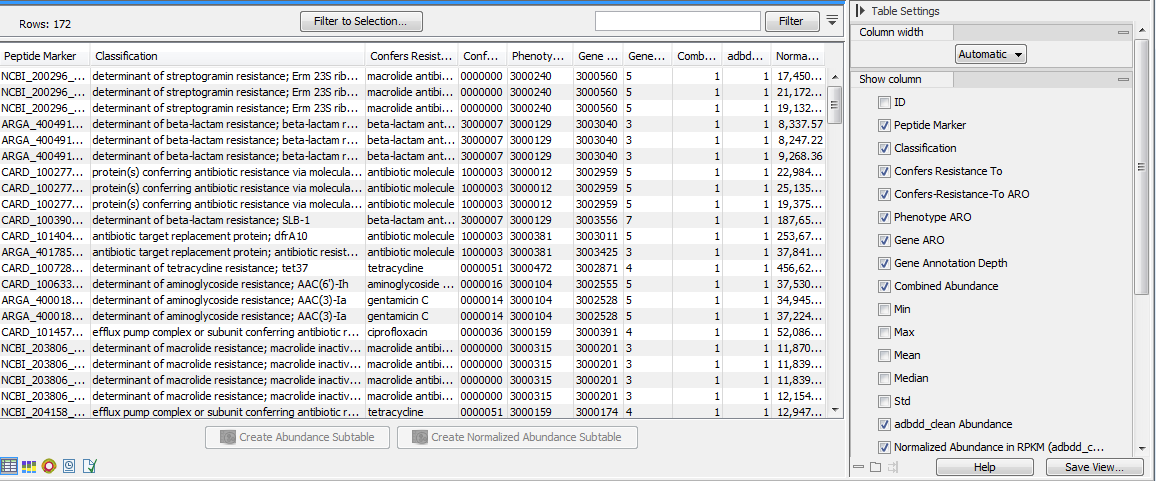
Genes | Free Full-Text | Metagenomic Information Recovery from Human Stool Samples Is Influenced by Sequencing Depth and Profiling Method
GitHub - borenstein-lab/MUSiCC: MUSiCC: A marker genes based framework for metagenomic normalization and accurate profiling of gene abundances in the microbiome

Association of Flavonifractor plautii, a Flavonoid-Degrading Bacterium, with the Gut Microbiome of Colorectal Cancer Patients in India | mSystems

Gene Expression Changes and Community Turnover Differentially Shape the Global Ocean Metatranscriptome - ScienceDirect